March 22, 2013
Well, I've started testing the predictions I made in my previous post, and exactly as I expected and deserved, I am getting killed. The actual results pretty consistently show more species than I predicted, sometimes way more.
I have started the process of debugging the problem. Of course, now that I have looked at the right answers, I can no longer use this data set for validation, especially since I plan to bang on my algorithm until it produces the right answers.
But in the interest of transparent science, I will at least document the debugging process. My first step was to review the most recent (and least-tested) code for obvious bugs, and I found one. I made an error parsing one of the data files, which had the effect of double-counting the total reads for each subject. I fixed that, but it didn't help the results much.
To start the debugging process, I am looking at the various places where my predictions could go wrong:
0) The data could be wrong. In particular, I assume that the rarefacted data I got from one data file is consistent with the complete dataset I got from another.
1) The posterior distribution could be right, but the predictive distribution could be wrong.
2) The posterior distribution might be wrong because of modeling errors.
3) The posterior distribution might be wrong because of implementation errors.
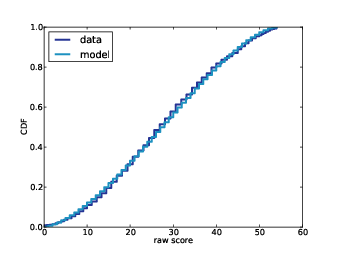
To check (0), I used the complete dataset to generate a few re-rarefacted datasets to see if my rarefaction process looks like theirs. It does, so I accept the data, at least for now.
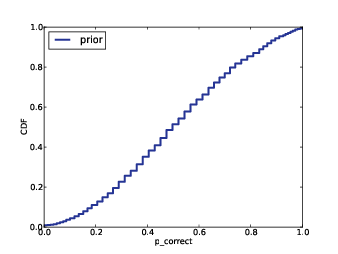
To check (1), I used the posterior distribution to generate a 90% credible interval for the total number of species. Since the number of observed species in the complete dataset is necessarily less than the total number of species, the actual values should fall in or below the CIs, but in fact they often exceed the CIs, meaning that there are just more species than my algorithm expects.
While investigating (1) I discovered one problem. In my prior distribution on the number of species, I was setting the upper bound too low, cutting off some values of n with non-negligible probability. So I cranked it up high enough that any additional increase has no further effect on the results. That helps, but my predictions are still too low.
The next step is to test (2). I will generate simulated datasets, generate predictions and then validation them. Since the simulated data come straight from the model, there can be no modeling errors. If I can't validate on simulated data, the problem has to be the algorithm or the implementation, not the model.
Of course, I should have done all this first, before blowing my testing data.
I won't have a chance to get back to this for a little while, but I'll update this post when I do.
Friday, March 22, 2013
Monday, February 18, 2013
Belly Button Biodiversity: Part Three
This is part three of a series of articles about a Bayesian solution to the Unseen Species problem, applied to data from the Belly Button Biodiversity project.
In Part One I presented the simplest version of the algorithm, which I think is easy to understand, but slow. In Think Bayes I present some ways to optimize it. In Part Two I apply the algorithm to real data and generate predictive distributions. Now in Part Three, as promised, I publish the predictions the algorithm generates. In Part Four I will compare the predictions to actual data.
Background: Belly Button Biodiversity 2.0 (BBB2) is a nation-wide citizen science project with the goal of identifying bacterial species that can be found in human navels (http://bbdata.yourwildlife.org).
Toward that end, I will now subject myself to public humiliation by generating a set of predictions using my almost-entirely-unvalidated solution to the Unseen Species problem. In the next installment I will publish the correct answers and score my predictions. Here are the details:
1) I am working with data from the Belly Button Biodiversity project; this data was used in a paper published in PLOS ONE and made available on the web pages of the journal and the researchers. The data consists of rDNA "reads" from 60 subjects. In order to facilitate comparisons between subjects, the researchers chose subjects with at least 400 reads, and for each subject they chose a random subset of 400 reads. The data for the other reads was not published.
2) For each subject, I know the results of the 400 selected reads, and the total number of reads. I will use my algorithm to generate a "prediction" for each subject, which is the number of additional species in the complete dataset.
3) Specifically, for each subject I will generate 9 posterior credible intervals (CIs) for the number of additional species: a 10% CI, a 20% CI, and so on up to a 90% CI.
4) To validate my predictions, I will count the number of CIs that contain the actual, correct value. Ideally, 10% of the correct values should fall in the 10% CIs, 20% should fall in the 20% CIs, and so on. Since the predictions and actual values are integers, a value that hits one end of a predicted CI counts as a half-hit.
Reading the last row, subject B975 yielded 1542 reads; in a random subset of 400 reads, there were 63 different species (or more precisely, OTUs). My algorithm predicts that if we look at all 1542 reads, the number of additional species we'll find is between 3 and 14, with 90% confidence.
I have to say that this table fills me with dread. The intervals seem quite small, which is to say that the algorithm is more confident than I am. The 90% CIs seem especially narrow to me; it is hard for me to believe that 90% of them will contain the correct values. Well, I guess that's why Karl Popper called them "bold hypotheses". We'll find out soon whether they are bold, or just reckless.
I want to thank Rob Dunn at BBB2 for his help with this project. The code and data I used to generate these results are available from this SVN repository.
EDIT 2-22-13: I ran the predictions again with more simulations. The results are not substantially different. I still haven't looked at the answers.
In Part One I presented the simplest version of the algorithm, which I think is easy to understand, but slow. In Think Bayes I present some ways to optimize it. In Part Two I apply the algorithm to real data and generate predictive distributions. Now in Part Three, as promised, I publish the predictions the algorithm generates. In Part Four I will compare the predictions to actual data.
Background: Belly Button Biodiversity 2.0 (BBB2) is a nation-wide citizen science project with the goal of identifying bacterial species that can be found in human navels (http://bbdata.yourwildlife.org).
Transparent science
In an effort to explore the limits of transparent science, I have started publishing my research in this blog as I go along. This past summer I wrote a series of articles exploring the relationship between Internet use and religious disaffiliation. This "publish as you go" model should help keep researchers honest. Among other things, it might mitigate publication bias due to the "file drawer effect." And if the data and code are published along with the results, that should help make experiments more reproducible.Toward that end, I will now subject myself to public humiliation by generating a set of predictions using my almost-entirely-unvalidated solution to the Unseen Species problem. In the next installment I will publish the correct answers and score my predictions. Here are the details:
1) I am working with data from the Belly Button Biodiversity project; this data was used in a paper published in PLOS ONE and made available on the web pages of the journal and the researchers. The data consists of rDNA "reads" from 60 subjects. In order to facilitate comparisons between subjects, the researchers chose subjects with at least 400 reads, and for each subject they chose a random subset of 400 reads. The data for the other reads was not published.
2) For each subject, I know the results of the 400 selected reads, and the total number of reads. I will use my algorithm to generate a "prediction" for each subject, which is the number of additional species in the complete dataset.
3) Specifically, for each subject I will generate 9 posterior credible intervals (CIs) for the number of additional species: a 10% CI, a 20% CI, and so on up to a 90% CI.
4) To validate my predictions, I will count the number of CIs that contain the actual, correct value. Ideally, 10% of the correct values should fall in the 10% CIs, 20% should fall in the 20% CIs, and so on. Since the predictions and actual values are integers, a value that hits one end of a predicted CI counts as a half-hit.
Predictions
And now, without further ado, here are my predictions. The columns labelled 10, 20, etc. are 10% credible intervals, 20% CIs, and so on.| Code | # reads | # species | 10 | 20 | 30 | 40 | 50 | 60 | 70 | 80 | 90 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| B1234 | 1392 | 48 | (4, 4) | (4, 5) | (3, 5) | (3, 5) | (3, 6) | (2, 6) | (2, 7) | (1, 7) | (1, 9) |
| B1235 | 2452 | 69 | (11, 12) | (10, 12) | (10, 13) | (9, 13) | (8, 14) | (8, 15) | (7, 16) | (6, 17) | (5, 19) |
| B1236 | 2964 | 45 | (4, 5) | (4, 5) | (4, 6) | (4, 6) | (3, 7) | (3, 7) | (3, 8) | (2, 9) | (1, 10) |
| B1237 | 3090 | 62 | (9, 10) | (9, 11) | (8, 11) | (8, 11) | (7, 12) | (7, 12) | (6, 13) | (5, 14) | (4, 16) |
| B1242 | 3056 | 61 | (9, 9) | (8, 10) | (8, 10) | (7, 11) | (7, 11) | (6, 12) | (6, 14) | (5, 15) | (5, 16) |
| B1243 | 1518 | 71 | (10, 11) | (10, 12) | (9, 12) | (8, 13) | (8, 13) | (8, 14) | (7, 15) | (6, 16) | (5, 17) |
| B1246 | 4230 | 91 | (23, 24) | (22, 25) | (21, 26) | (21, 27) | (19, 28) | (18, 29) | (17, 30) | (16, 33) | (14, 35) |
| B1253 | 1928 | 86 | (16, 17) | (15, 18) | (14, 18) | (14, 20) | (13, 20) | (13, 21) | (12, 23) | (11, 24) | (10, 26) |
| B1254 | 918 | 58 | (5, 5) | (4, 6) | (4, 6) | (3, 6) | (3, 7) | (3, 7) | (2, 8) | (2, 9) | (1, 10) |
| B1258 | 1350 | 87 | (15, 16) | (14, 17) | (14, 17) | (13, 18) | (12, 19) | (11, 19) | (11, 20) | (10, 21) | (8, 24) |
| B1259 | 1002 | 80 | (10, 11) | (10, 12) | (10, 12) | (9, 13) | (9, 14) | (8, 14) | (7, 15) | (6, 16) | (6, 18) |
| B1260 | 1944 | 96 | (22, 23) | (21, 24) | (20, 25) | (19, 25) | (19, 26) | (18, 27) | (17, 29) | (15, 30) | (14, 32) |
| B1264 | 1122 | 83 | (12, 13) | (12, 14) | (11, 14) | (10, 15) | (10, 15) | (9, 16) | (8, 17) | (7, 18) | (6, 20) |
| B1265 | 2928 | 85 | (18, 19) | (17, 20) | (16, 21) | (16, 22) | (15, 23) | (14, 24) | (13, 25) | (12, 26) | (11, 28) |
| B1273 | 2980 | 61 | (9, 9) | (8, 10) | (8, 10) | (7, 11) | (7, 12) | (6, 12) | (6, 13) | (5, 14) | (4, 16) |
| B1275 | 1672 | 85 | (16, 17) | (15, 18) | (15, 19) | (14, 19) | (13, 20) | (13, 21) | (12, 22) | (11, 24) | (9, 25) |
| B1278 | 1242 | 47 | (4, 4) | (3, 4) | (3, 5) | (3, 5) | (2, 6) | (2, 6) | (2, 6) | (2, 7) | (1, 8) |
| B1280 | 1772 | 46 | (4, 4) | (4, 5) | (3, 5) | (3, 5) | (3, 6) | (2, 6) | (2, 7) | (2, 8) | (1, 9) |
| B1282 | 1132 | 67 | (8, 9) | (7, 9) | (7, 10) | (6, 10) | (6, 11) | (6, 11) | (5, 12) | (5, 13) | (3, 15) |
| B1283 | 1414 | 67 | (8, 9) | (8, 10) | (7, 10) | (7, 11) | (7, 11) | (6, 12) | (5, 13) | (4, 14) | (3, 16) |
| B1284 | 1158 | 91 | (15, 16) | (14, 17) | (14, 17) | (13, 18) | (13, 19) | (12, 19) | (12, 20) | (10, 21) | (9, 23) |
| B1285 | 2340 | 55 | (7, 7) | (6, 8) | (6, 8) | (5, 8) | (5, 9) | (4, 9) | (4, 10) | (3, 12) | (2, 13) |
| B1286 | 1728 | 66 | (9, 10) | (9, 11) | (8, 11) | (8, 12) | (8, 12) | (7, 13) | (6, 14) | (6, 14) | (4, 16) |
| B1288 | 1280 | 107 | (23, 24) | (22, 25) | (21, 25) | (21, 26) | (20, 27) | (19, 27) | (18, 29) | (17, 31) | (15, 32) |
| B1289 | 2054 | 103 | (26, 27) | (25, 28) | (24, 29) | (23, 30) | (23, 30) | (22, 32) | (21, 33) | (20, 34) | (17, 36) |
| B1291 | 1248 | 94 | (17, 18) | (16, 19) | (16, 20) | (15, 20) | (15, 21) | (13, 22) | (13, 23) | (12, 25) | (10, 27) |
| B1292 | 1864 | 82 | (15, 16) | (14, 16) | (13, 17) | (13, 18) | (13, 19) | (12, 20) | (11, 21) | (10, 22) | (9, 24) |
| B1293 | 1904 | 76 | (13, 14) | (12, 14) | (12, 15) | (11, 16) | (11, 16) | (10, 17) | (9, 18) | (8, 19) | (7, 22) |
| B1294 | 1784 | 78 | (14, 15) | (13, 16) | (12, 16) | (12, 17) | (11, 18) | (11, 19) | (10, 19) | (9, 20) | (8, 23) |
| B1295 | 1408 | 70 | (10, 10) | (9, 11) | (9, 12) | (8, 12) | (8, 12) | (7, 13) | (7, 14) | (6, 15) | (4, 17) |
| B1296 | 2034 | 55 | (7, 7) | (6, 8) | (6, 8) | (6, 8) | (5, 9) | (4, 9) | (4, 10) | (4, 11) | (3, 12) |
| B1298 | 1478 | 72 | (10, 11) | (9, 12) | (9, 12) | (9, 13) | (8, 13) | (8, 14) | (7, 15) | (6, 16) | (5, 18) |
| B1308 | 1160 | 58 | (6, 6) | (5, 7) | (5, 7) | (5, 7) | (4, 8) | (4, 8) | (3, 9) | (3, 10) | (2, 11) |
| B1310 | 1066 | 80 | (11, 12) | (11, 13) | (10, 13) | (9, 14) | (9, 15) | (8, 15) | (7, 16) | (7, 17) | (5, 19) |
| B1374 | 2364 | 48 | (5, 5) | (4, 6) | (4, 6) | (4, 6) | (3, 7) | (3, 7) | (3, 8) | (2, 9) | (2, 10) |
| B940 | 2874 | 93 | (22, 24) | (21, 25) | (21, 25) | (20, 26) | (19, 27) | (19, 28) | (18, 30) | (16, 32) | (14, 33) |
| B941 | 2154 | 48 | (5, 6) | (5, 6) | (4, 6) | (4, 7) | (4, 7) | (3, 7) | (3, 8) | (2, 9) | (2, 11) |
| B944 | 954 | 52 | (4, 4) | (4, 5) | (3, 5) | (3, 5) | (3, 6) | (2, 6) | (2, 6) | (2, 7) | (1, 9) |
| B945 | 2390 | 67 | (10, 11) | (10, 12) | (9, 12) | (9, 13) | (8, 13) | (8, 14) | (7, 15) | (7, 16) | (5, 17) |
| B946 | 5012 | 85 | (20, 21) | (19, 22) | (19, 23) | (18, 24) | (18, 24) | (17, 26) | (16, 27) | (15, 28) | (12, 31) |
| B947 | 3356 | 62 | (10, 11) | (9, 11) | (9, 12) | (8, 12) | (7, 13) | (7, 14) | (6, 14) | (5, 15) | (5, 17) |
| B948 | 2384 | 80 | (16, 17) | (15, 18) | (14, 18) | (14, 19) | (13, 20) | (12, 21) | (11, 22) | (10, 23) | (9, 25) |
| B950 | 1560 | 63 | (8, 9) | (8, 10) | (8, 10) | (7, 10) | (7, 11) | (6, 11) | (5, 12) | (5, 13) | (4, 15) |
| B952 | 1648 | 57 | (7, 7) | (6, 8) | (6, 8) | (6, 8) | (5, 9) | (5, 9) | (4, 10) | (3, 11) | (3, 12) |
| B953 | 1452 | 32 | (2, 2) | (1, 2) | (1, 2) | (1, 3) | (1, 3) | (1, 3) | (0, 3) | (0, 4) | (0, 5) |
| B954 | 1996 | 29 | (2, 2) | (1, 2) | (1, 2) | (1, 2) | (1, 3) | (1, 3) | (0, 3) | (0, 4) | (0, 4) |
| B955 | 1474 | 65 | (8, 9) | (8, 9) | (7, 9) | (7, 10) | (7, 10) | (6, 11) | (5, 12) | (5, 13) | (4, 14) |
| B956 | 1482 | 71 | (10, 11) | (10, 12) | (10, 12) | (9, 13) | (8, 14) | (8, 14) | (7, 15) | (6, 16) | (5, 18) |
| B957 | 2604 | 36 | (3, 3) | (3, 3) | (2, 4) | (2, 4) | (2, 5) | (1, 5) | (1, 6) | (1, 6) | (1, 7) |
| B958 | 2840 | 29 | (2, 2) | (2, 2) | (1, 2) | (1, 3) | (1, 3) | (1, 3) | (1, 4) | (0, 4) | (0, 5) |
| B961 | 1214 | 36 | (2, 3) | (2, 3) | (2, 3) | (2, 4) | (1, 4) | (1, 4) | (1, 5) | (1, 5) | (0, 6) |
| B962 | 1138 | 41 | (3, 3) | (2, 3) | (2, 3) | (2, 4) | (2, 4) | (1, 4) | (1, 5) | (1, 6) | (0, 7) |
| B963 | 1600 | 71 | (10, 12) | (10, 12) | (9, 12) | (9, 13) | (8, 14) | (8, 14) | (7, 15) | (5, 16) | (4, 19) |
| B966 | 1950 | 80 | (15, 16) | (14, 16) | (14, 17) | (13, 17) | (12, 18) | (11, 18) | (11, 20) | (10, 22) | (9, 23) |
| B967 | 1108 | 47 | (3, 4) | (3, 4) | (3, 4) | (2, 5) | (2, 5) | (2, 5) | (2, 6) | (1, 7) | (1, 7) |
| B968 | 2432 | 52 | (6, 7) | (6, 7) | (6, 7) | (5, 8) | (5, 8) | (4, 9) | (3, 9) | (3, 10) | (2, 11) |
| B971 | 1462 | 49 | (4, 5) | (4, 5) | (4, 5) | (3, 6) | (3, 6) | (3, 7) | (2, 7) | (2, 8) | (1, 9) |
| B972 | 1438 | 75 | (11, 12) | (11, 13) | (10, 13) | (10, 14) | (9, 14) | (8, 15) | (8, 16) | (7, 17) | (6, 18) |
| B974 | 5072 | 54 | (7, 8) | (7, 9) | (6, 9) | (6, 9) | (5, 10) | (5, 10) | (4, 11) | (4, 12) | (2, 14) |
| B975 | 1542 | 63 | (7, 8) | (7, 9) | (6, 9) | (6, 9) | (6, 10) | (5, 11) | (5, 11) | (4, 12) | (3, 14) |
Reading the last row, subject B975 yielded 1542 reads; in a random subset of 400 reads, there were 63 different species (or more precisely, OTUs). My algorithm predicts that if we look at all 1542 reads, the number of additional species we'll find is between 3 and 14, with 90% confidence.
I have to say that this table fills me with dread. The intervals seem quite small, which is to say that the algorithm is more confident than I am. The 90% CIs seem especially narrow to me; it is hard for me to believe that 90% of them will contain the correct values. Well, I guess that's why Karl Popper called them "bold hypotheses". We'll find out soon whether they are bold, or just reckless.
I want to thank Rob Dunn at BBB2 for his help with this project. The code and data I used to generate these results are available from this SVN repository.
EDIT 2-22-13: I ran the predictions again with more simulations. The results are not substantially different. I still haven't looked at the answers.
Friday, February 8, 2013
Belly Button Biodiversity: Part Two
This is part two of a series of articles about a Bayesian solution to the Unseen Species problem, applied to data from the Belly Button Biodiversity project.
In Part One I presented the simplest version of the algorithm, which I think is easy to understand, but slow. In Think Bayes I present some ways to optimize it. Now in Part Two I apply the algorithm to real data and generate predictive distributions. In Part Three I will publish the predictions the algorithm generates, and in Part Four I will compare the predictions to actual data.
Background: Belly Button Biodiversity 2.0 (BBB2) is a nation-wide citizen science project with the goal of identifying bacterial species that can be found in human navels (http://bbdata.yourwildlife.org).
The belly button data
To get a sense of what the data look like, consider subject B1242, whose sample of 400 reads yielded 61 species with the following counts:
92, 53, 47, 38, 15, 14, 12, 10, 8, 7, 7, 5, 5, 4, 4, 4, 4, 4, 4, 4, 3, 3, 3, 3, 3, 3, 3, 2, 2, 2, 2, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1There are a few dominant species that make up a substantial fraction of the whole, but many species that yielded only a single read. The number of these “singletons” suggests that there are likely to be at least a few unseen species.
In the example with lions and tigers, we assume that each animal in the preserve is equally likely to be observed. Similarly, for the belly button data, we assume that each bacterium is equally likely to yield a read.
In reality, it is possible that each step in the data-collection process might introduce consistent biases. Some species might be more likely to be picked up by a swab, or to yield identifiable amplicons. So when we talk about the prevalence of each species, we should remember this source of error.
I should also acknowledge that I am using the term “species” loosely. First, bacterial species are not well-defined. Second, some reads identify a particular species, others only identify a genus. To be more precise, I should say “operational taxonomic unit”, or OTU.
Now let’s process some of the belly button data. I defined a class called Subject to represent information about each subject in the study:
class Subject(object):
def __init__(self, code):
self.code = code
self.species = []
Each subject has a string code, like “B1242”, and a list of (count, species name) pairs, sorted in increasing order by count. Subject provides several methods to make it easy to these counts and species names. You can see the details in http://thinkbayes.com/species.py.In addition, Subject.Process creates a suite, specifically a suite of type Species5, which represents the distribution of n and the prevalences after processing the data.
Figure 12.3: Distribution of n for subject B1242.
It also provides PlotDistOfN, which plots the posterior distribution of n. Figure 12.3 shows this distribution for subject B1242. The probability that there are exactly 61 species, and no unseen species, is nearly zero. The most likely value is 72, with 90% credible interval 66 to 79. At the high end, it is unlikely that there are as many as 87 species.
Next we compute the posterior distribution of prevalence for each species. Species2 providesDistOfPrevalence:
# class Species2
def DistOfPrevalence(self, index):
pmfs = thinkbayes.Pmf()
for n, prob in zip(self.ns, self.probs):
beta = self.MarginalBeta(n, index)
pmf = beta.MakePmf()
pmfs.Set(pmf, prob)
mix = thinkbayes.MakeMixture(pmfs)
return pmfs, mix
index indicates which species we want. For each value of n, we have a different posterior distribution of prevalence.So the loop iterates through the possible values of n and their probabilities. For each value of n it gets a Beta object representing the marginal distribution for the indicated species. Remember that Beta objects contain the parameters alpha and beta; they don’t have values and probabilities like a Pmf, but they provide MakePmf which generates a discrete approximation to the continuous beta distribution.
Figure 12.4: Distribution of prevalences for subject B1242.
pmfs is a MetaPmf that contains the distributions of prevalence, conditioned on n. MakeMixturecombines the MetaPmf into mix, which combines the conditional distributions into the answer, a single distribution of prevalence.
Figure 12.4 shows these distributions for the five species with the most reads. The most prevalent species accounts for 23% of the 400 reads, but since there are almost certainly unseen species, the most likely estimate for its prevalence is 20%, with 90% credible interval between 17% and 23%.
Predictive distributions
I introduced the hidden species problem in the form of four related questions. We have answered the first two by computing the posterior distribution for n and the prevalence of each species.
Figure 12.5: Simulated rarefaction curves for subject B1242.
The other two questions are:
- If we are planning to collect additional samples, can we predict how many new species we are likely to discover?
- How many additional reads are needed to increase the fraction of observed species to a given threshold?
The kernel of these simulations looks like this:
- Choose n from its posterior distribution.
- Choose a prevalence for each species, including possible unseen species, using the Dirichlet distribution.
- Generate a random sequence of future observations.
- Compute the number of new species,
num_new, as a function of the number of additional samples, k. - Repeat the previous steps and accumulate the joint distribution of
num_newand k.
# class Subject
def RunSimulation(self, num_samples):
m, seen = self.GetSeenSpecies()
n, observations = self.GenerateObservations(num_samples)
curve = []
for k, obs in enumerate(observations):
seen.add(obs)
num_new = len(seen) - m
curve.append((k+1, num_new))
return curve
num_samples is the number of additional samples to simulate. m is the number of seen species, and seen is a set of strings with a unique name for each species. n is a random value from the posterior distribution, and observations is a random sequence of species names.
The result of RunSimulation is a “rarefaction curve”, represented as a list of pairs with the number of samples and the number of new species seen.
Before we see the results, let’s look at GetSeenSpecies and GenerateObservations.
#class Subject
def GetSeenSpecies(self):
names = self.GetNames()
m = len(names)
seen = set(SpeciesGenerator(names, m))
return m, seen
GetNames returns the list of species names that appear in the data files, but for many subjects these names are not unique. So I use SpeciesGenerator to extend each name with a serial number:def SpeciesGenerator(names, num):
i = 0
for name in names:
yield '%s-%d' % (name, i)
i += 1
while i < num:
yield 'unseen-%d' % i
i += 1
Given a name like Corynebacterium, SpeciesGenerator yields Corynebacterium-1. When the list of names is exhausted, it yields names like unseen-62.
Here is GenerateObservations:
# class Subject
def GenerateObservations(self, num_samples):
n, prevalences = self.suite.Sample()
names = self.GetNames()
name_iter = SpeciesGenerator(names, n)
d = dict(zip(name_iter, prevalences))
cdf = thinkbayes.MakeCdfFromDict(d)
observations = cdf.Sample(num_samples)
return n, observations
Again, num_samples is the number of additional samples to generate. n and prevalences are samples from the posterior distribution.
cdf is a Cdf object that maps species names, including the unseen, to cumulative probabilities. Using a Cdf makes it efficient to generate a random sequence of species names.
Finally, here is Species2.Sample:
def Sample(self):
pmf = self.DistOfN()
n = pmf.Random()
prevalences = self.SampleConditional(n)
return n, prevalences
And SampleConditional, which generates a sample of prevalences conditioned on n:# class Species2
def SampleConditional(self, n):
params = self.params[:n]
gammas = numpy.random.gamma(params)
gammas /= gammas.sum()
return gammas
We saw this algorithm for generating prevalences previously in Species2.SampleLikelihood.
Figure 12.5 shows 100 simulated rarefaction curves for subject B1242. I shifted each curve by a random offset so they would not all overlap. By inspection we can estimate that after 400 more samples we are likely to find 2–6 new species.
Joint posterior
To be more precise, we can use the simulations to estimate the joint distribution of
Figure 12.6: Distributions of the number of new species conditioned on the number of additional samples.
num_new andk, and from that we can get the distribution of num_new conditioned on any value of k.# class Subject
def MakeJointPredictive(self, curves):
joint = thinkbayes.Joint()
for curve in curves:
for k, num_new in curve:
joint.Incr((k, num_new))
joint.Normalize()
return joint
MakeJointPredictive makes a Joint object, which is a Pmf whose values are tuples.
curves is a list of rarefaction curves created by RunSimulation. Each curve contains a list of pairs of k and
num_new.
The resulting joint distribution is a map from each pair to its probability of occurring. Given the joint distribution, we can get the distribution of
num_new conditioned on k:# class Joint
def Conditional(self, i, j, val):
pmf = Pmf()
for vs, prob in self.Items():
if vs[j] != val: continue
pmf.Incr(vs[i], prob)
pmf.Normalize()
return pmf
i is the index of the variable whose distribution we want; j is the index of the conditional variables, and val is the value the jth variable has to have. You can think of this operation as taking vertical slices out of Figure 12.5.
Subject.MakeConditionals takes a list of ks and computes the conditional distribution of
num_new for each k. The result is a list of Cdf objects.# class Subject
def MakeConditionals(self, curves, ks):
joint = self.MakeJointPredictive(curves)
cdfs = []
for k in ks:
pmf = joint.Conditional(1, 0, k)
pmf.name = 'k=%d' % k
cdf = thinkbayes.MakeCdfFromPmf(pmf)
cdfs.append(cdf)
return cdfs
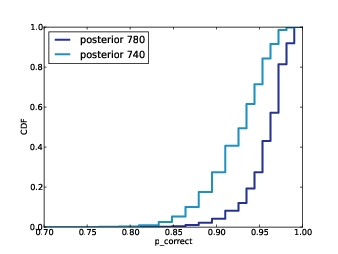
Figure 12.6 shows the results. After 100 samples, the median predicted number of new species is 2; the 90% credible interval is 0 to 5. After 800 samples, we expect to see 3 to 12 new species.Coverage
The last question we want to answer is, “How many additional reads are needed to increase the fraction of observed species to a given threshold?”
Figure 12.7: Complementary CDF of coverage for a range of additional samples.
To answer this question, we’ll need a version of RunSimulation that computes the fraction of observed species rather than the number of new species.
# class Subject
def RunSimulation(self, num_samples):
m, seen = self.GetSeenSpecies()
n, observations = self.GenerateObservations(num_samples)
curve = []
for k, obs in enumerate(observations):
seen.add(obs)
frac_seen = len(seen) / float(n)
curve.append((k+1, frac_seen))
return curve
Next we loop through each curve and make a dictionary, d, that maps from the number of additional samples, k, to a list of fracs; that is, a list of values for the coverage achieved after ksamples. def MakeFracCdfs(self, curves):
d = {}
for curve in curves:
for k, frac in curve:
d.setdefault(k, []).append(frac)
cdfs = {}
for k, fracs in d.iteritems():
cdf = thinkbayes.MakeCdfFromList(fracs)
cdfs[k] = cdf
return cdfs
Then for each value of k we make a Cdf of fracs; this Cdf represents the distribution of coverage after k samples.
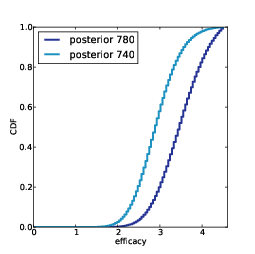
Remember that the CDF tells you the probability of falling below a given threshold, so thecomplementary CDF tells you the probability of exceeding it. Figure 12.7 shows complementary CDFs for a range of values of k.
To read this figure, select the level of coverage you want to achieve along the x-axis. As an example, choose 90%.
Now you can read up the chart to find the probability of achieving 90% coverage after k samples. For example, with 300 samples, you have about a 60% of getting 90% coverage. With 700 samples, you have a 90% chance of getting 90% coverage.
With that, we have answered the four questions that make up the unseen species problem. Next time: validation!
Tuesday, February 5, 2013
Belly Button Biodiversity: Part One
This post is a excerpt from Think Bayes: Bayesian Statistics Made Simple, the book I am working on now. You can read the entire current draft at http://thinkbayes.com.
The project might seem whimsical, but it is part of an increasing interest in the human microbiome, the set of microorganisms that live on human skin and other surfaces that contact the environment.
In their pilot study, BBB2 researchers collected swabs from the navels of 60 volunteers, used multiplex pyrosequencing to extract and sequence fragments of 16S rDNA, then identified the species or genus the fragments came from. Each identified fragment is called a “read.”
We can use these data to answer several related questions:
If we have an equal chance of observing any animal in the preserve then the number of each species we see is governed by the multinomial distribution. If the prevalence of lions and tigers and bears is
But there are two problems:
The Dirichlet distribution is the multi-dimensional generalization of the beta distribution. Instead of two possible outcomes, like heads and tails, the Dirichlet distribution handles any number of outcomes: in this example, three species.
If there are n outcomes, the Dirichlet distribution is described by n parameters, written αi.
Here’s the definition, from thinkbayes.py, of a class that represents a Dirichlet distribution:
Given a Dirichlet distribution, the marginal distribution for each prevalence is a beta distribution, which we can compute like this:
In the example, the prior marginal distribution for each species is Beta(1, 2). We can compute the prior means like this:
To update the Dirichlet distribution, we add the number of observations to each parameter, like this:
But data can be shorter than params; in that case there are some hypothetical species that have not been observed.
Here’s code that updates dirichlet with the observed data and computes the posterior marginal distributions.

Now let’s get back to the original problem, estimating the total number of species. To solve this problem I’ll define a metasuite, which is a Suite that contains other Suites as hypotheses. In this case, the top-level Suite contains hypotheses about the number of species; the bottom level contains hypotheses about prevalences. A multi-level model like this is called “hierarchical.”
Here’s the class definition:
Here’s the code that creates the top-level suite:
To update a hierarchical model, you have to update all levels. Sometimes it is necessary or more efficient to update the bottom level first and work up. In this case it doesn’t matter, so I update the top level first:
Now all we need is a likelihood function. As usual, Likelihood gets a hypothesis and a dataset as arguments:
Why do we have to call it 1000 times? Because Dirichlet.Likelihood doesn’t actually compute the likelihood of the data under the whole Dirichlet distribution. Instead, it draws one sample from the hypothetical distribution and computes the likelihood of the data under the sampled set of prevalences.
Here’s what it looks like:
Otherwise we select a random set of prevalences, p, and compute the multinomial PDF, which is
pi is the prevalence of the ith species, and xi is the
observed number. The first term, c(x), is the multinomial
coefficient; I left it out of the computation because it is
a multiplicative factor that depends only
on the data, not the hypothesis, so it gets normalized away
(see http://en.wikipedia.org/wiki/Multinomial_distribution).
Also, I truncated p at m, which is the number of observed species. For the unseen species, xi is 0, so pixi is 1, so we can leave them out of the product.
A less obvious, but faster, way is to select values from n gamma distributions, then normalize by dividing through by the total. Here’s the code:
The most likely value is 5. Values from 3 to 8 are all likely; after that the probabilities drop off quickly. The probability that there are 29 species is low enough to be negligible; if we chose a higher bound, we would get the same result.
But remember that we started with a uniform prior for n. If we have background information about the number of species in the environment, we might choose a different prior.
The only problem is that it is slow. It’s good enough for the example with only 3 observed species, but not good enough for the belly button data, with more than 100 species in some samples.
In Think Bayes I present a series of optimizations we need to make this solution scale. Here’s a road map of the steps:
Belly button bacteria
Belly Button Biodiversity 2.0 (BBB2) is a nation-wide citizen science project with the goal of identifying bacterial species that can be found in human navels (http://bbdata.yourwildlife.org).
The project might seem whimsical, but it is part of an increasing interest in the human microbiome, the set of microorganisms that live on human skin and other surfaces that contact the environment.
In their pilot study, BBB2 researchers collected swabs from the navels of 60 volunteers, used multiplex pyrosequencing to extract and sequence fragments of 16S rDNA, then identified the species or genus the fragments came from. Each identified fragment is called a “read.”
We can use these data to answer several related questions:
- Based on the number of species observed, can we estimate the total number of species in the environment?
- Can we estimate the prevalence of each species; that is, the fraction of the total population belonging to each species?
- If we are planning to collect additional samples, can we predict how many new species we are likely to discover?
- How many additional reads are needed to increase the fraction of observed species to a given threshold?
Lions and tigers and bears
I’ll start with a simplified version of the problem where we know that there are exactly three species. Let’s call them lions, tigers and bears. Suppose we visit a wild animal preserve and see 3 lions, 2 tigers and one bear.
If we have an equal chance of observing any animal in the preserve then the number of each species we see is governed by the multinomial distribution. If the prevalence of lions and tigers and bears is
p_lion and p_tiger and p_bear, the likelihood of
seeing 3 lions, 2 tigers and one bear isp_lion**3 * p_tiger**2 * p_bear**1
An approach that is tempting, but not correct, is to use beta
distributions, as in Section 4.6, to describe the prevalence of
each species separately. For example, we saw 3 lions and 3 non-lions;
if we think of that as 3 “heads” and 3 “tails,” then the posterior
distribution of p_lion is: beta = thinkbayes.Beta()
beta.Update((3, 3))
print beta.MaximumLikelihood()
The maximum likelihood estimate for
p_lion is the observed
rate, 50%. Similarly the MLEs for p_tiger and p_bear
are 33% and 17%.But there are two problems:
- We have implicitly used a prior for each species that is uniform from 0 to 1, but since we know that there are three species, that prior is not correct. The right prior should have a mean of 1/3, and there should be zero likelihood that any species has a prevalence of 100%.
- The distributions for each species are not independent, because the prevalences have to add up to 1. To capture this dependence, we need a joint distribution for the three prevalences.
p_lion,
p_tiger and p_bear.The Dirichlet distribution is the multi-dimensional generalization of the beta distribution. Instead of two possible outcomes, like heads and tails, the Dirichlet distribution handles any number of outcomes: in this example, three species.
If there are n outcomes, the Dirichlet distribution is described by n parameters, written αi.
Here’s the definition, from thinkbayes.py, of a class that represents a Dirichlet distribution:
class Dirichlet(object):
def __init__(self, n):
self.n = n
self.params = numpy.ones(n, dtype=numpy.int)
n is the number of dimensions; initially the parameters are all 1. I use a numpy array to store the parameters so I can take advantage of array operations.
Given a Dirichlet distribution, the marginal distribution for each prevalence is a beta distribution, which we can compute like this:
def MarginalBeta(self, i):
alpha0 = self.params.sum()
alpha = self.params[i]
return Beta(alpha, alpha0-alpha)
i is the index of the marginal distribution we want. alpha0 is the sum of the parameters; alpha is the parameter for the given species.
In the example, the prior marginal distribution for each species is Beta(1, 2). We can compute the prior means like this:
dirichlet = thinkbayes.Dirichlet(3)
for i in range(3):
beta = dirichlet.MarginalBeta(i)
print beta.Mean()
As expected, the prior mean prevalence for each species is 1/3.
To update the Dirichlet distribution, we add the number of observations to each parameter, like this:
def Update(self, data):
m = len(data)
self.params[:m] += data
Here data is a sequence of counts in the same order as params, so in this example, it should be the number of lions, tigers and bears.
But data can be shorter than params; in that case there are some hypothetical species that have not been observed.
Here’s code that updates dirichlet with the observed data and computes the posterior marginal distributions.
data = [3, 2, 1]
dirichlet.Update(data)
for i in range(3):
beta = dirichlet.MarginalBeta(i)
pmf = beta.MakePmf()
print i, pmf.Mean()
This figure shows the results. The posterior mean prevalences are 44%, 33% and 22%.

A hierarchical model
We have solved a simplified version of the problem: if we know how many species there are, we can estimate the prevalence of each.
Now let’s get back to the original problem, estimating the total number of species. To solve this problem I’ll define a metasuite, which is a Suite that contains other Suites as hypotheses. In this case, the top-level Suite contains hypotheses about the number of species; the bottom level contains hypotheses about prevalences. A multi-level model like this is called “hierarchical.”
Here’s the class definition:
class Species(thinkbayes.Suite):
def __init__(self, ns):
hypos = [thinkbayes.Dirichlet(n) for n in ns]
thinkbayes.Suite.__init__(self, hypos)
__init__ takes a list of possible values for n and
makes a list of Dirichlet objects.Here’s the code that creates the top-level suite:
ns = range(3, 30)
suite = Species(ns)
ns is the list of possible values for n. We have seen 3 species, so there have to be at least that many. I chose an upper bound that seemed reasonable, but we will have to check later that the probability of exceeding this bound is low. And at least initially we assume that any value in this range is equally likely.
To update a hierarchical model, you have to update all levels. Sometimes it is necessary or more efficient to update the bottom level first and work up. In this case it doesn’t matter, so I update the top level first:
#class Species
def Update(self, data):
thinkbayes.Suite.Update(self, data)
for hypo in self.Values():
hypo.Update(data)
Species.Update invokes Update in the parent class, then loops through the sub-hypotheses and updates them.
Now all we need is a likelihood function. As usual, Likelihood gets a hypothesis and a dataset as arguments:
# class Species
def Likelihood(self, hypo, data):
dirichlet = hypo
like = 0
for i in range(1000):
like += dirichlet.Likelihood(data)
return like
hypo is a Dirichlet object; data is a sequence of observed counts. Species.Likelihood calls Dirichlet.Likelihood 1000 times and returns the total.
Why do we have to call it 1000 times? Because Dirichlet.Likelihood doesn’t actually compute the likelihood of the data under the whole Dirichlet distribution. Instead, it draws one sample from the hypothetical distribution and computes the likelihood of the data under the sampled set of prevalences.
Here’s what it looks like:
# class Dirichlet
def Likelihood(self, data):
m = len(data)
if self.n < m:
return 0
x = data
p = self.Random()
q = p[:m]**x
return q.prod()
The length of data is the number of species observed. If we see more species than we thought existed, the likelihood is 0.
Otherwise we select a random set of prevalences, p, and compute the multinomial PDF, which is
| c(x) |
| pixi |
Also, I truncated p at m, which is the number of observed species. For the unseen species, xi is 0, so pixi is 1, so we can leave them out of the product.
Random sampling
There are two ways to generate a random sample from a Dirichlet distribution. One is to use the marginal beta distributions, but in that case you have to select one at a time and scale the rest so they add up to 1 (see http://en.wikipedia.org/wiki/Dirichlet_distribution#Random_number_generation).
A less obvious, but faster, way is to select values from n gamma distributions, then normalize by dividing through by the total. Here’s the code:
# class Dirichlet
def Random(self):
p = numpy.random.gamma(self.params)
return p / p.sum()
Now we’re ready to look at some results. Here is the code that updates the top-level suite and extracts the posterior PMF of n:
data = [3, 2, 1]
suite.Update(data)
pmf = suite.DistOfN()
To get the posterior distribution of n, DistOfN iterates through the top-level hypotheses:
def DistOfN(self):
pmf = thinkbayes.Pmf()
for hypo, prob in self.Items():
pmf.Set(hypo.n, prob)
return pmf
This figure shows the result:
The most likely value is 5. Values from 3 to 8 are all likely; after that the probabilities drop off quickly. The probability that there are 29 species is low enough to be negligible; if we chose a higher bound, we would get the same result.
But remember that we started with a uniform prior for n. If we have background information about the number of species in the environment, we might choose a different prior.
Optimization
I have to admit that I am proud of this example. The unseen species problem is not easy, and I think this solution is simple and clear, and takes surprisingly few lines of code (about 50 so far).
The only problem is that it is slow. It’s good enough for the example with only 3 observed species, but not good enough for the belly button data, with more than 100 species in some samples.
In Think Bayes I present a series of optimizations we need to make this solution scale. Here’s a road map of the steps:
- The first step is to recognize that if we update the Dirichlet distributions with the same data, the first m parameters are the same for all of them. The only difference is the number of hypothetical unseen species. So we don’t really need n Dirichlet objects; we can store the parameters in the top level of the hierarchy. Species2 implements this optimization.
- Species2 also uses the same set of random values for all of the hypotheses. This saves time generating random values, but it has a second benefit that turns out to be more important: by giving all hypothesis the same selection from the sample space, we make the comparison between the hypotheses more fair, so it takes fewer iterations to converge.
- But there is still a major performance problem. As the number of observed species increases, the array of random prevalences gets bigger, and the chance of choosing one that is approximately right becomes small. So the vast majority of iterations yield small likelihoods that don’t contribute much to the total, and don’t discriminate between hypotheses.The solution is to do the updates one species at a time. Species4 is a simple implementation of this strategy using Dirichlet objects to represent the sub-hypotheses.
- Finally, Species5 combines the sub-hypothesis into the top level and uses numpy array operations to speed things up.
Wednesday, January 23, 2013
Bayesian Statistics Made Simple
I am happy to announce that I will offer an updated and revised version of my tutorial, Bayesian Statistics Made Simple, at PyCon 2013.
Registration is open now. Here are the details:
PyCon 2013
Santa Clara, CA
Wednesday 13 March, 1:20 p.m.–4:40 p.m.
Bayesian statistics made simple
Allen Downey
Audience level: Intermediate
DESCRIPTION
An introduction to Bayesian statistics using Python. Bayesian statistics are usually presented mathematically, but many of the ideas are easier to understand computationally. People who know some Python have a head start.
We will use material from Think Stats: Probability and Statistics for Programmers (O’Reilly Media), and Think Bayes, a work in progress at http://thinkbayes.com.
ABSTRACT
Bayesian statistical methods are becoming more common and more important, but there are not many resources to help beginners get started. People who know Python can use their programming skills to get a head start.
I will present simple programs that demonstrate the concepts of Bayesian statistics, and apply them to a range of example problems. Participants will work hands-on with example code and practice on example problems.
Students should have at least basic Python and basic statistics. If you learned about Bayes’s Theorem and probability distributions at some time, that’s enough, even if you don’t remember it!
Students should bring a laptop with Python 2.x and matplotlib. You can work in any environment; you just need to be able to download a Python program and run it.
Registration is open now. Here are the details:
PyCon 2013
Santa Clara, CA
Wednesday 13 March, 1:20 p.m.–4:40 p.m.
Bayesian statistics made simple
Allen Downey
Audience level: Intermediate
DESCRIPTION
An introduction to Bayesian statistics using Python. Bayesian statistics are usually presented mathematically, but many of the ideas are easier to understand computationally. People who know some Python have a head start.
We will use material from Think Stats: Probability and Statistics for Programmers (O’Reilly Media), and Think Bayes, a work in progress at http://thinkbayes.com.
ABSTRACT
Bayesian statistical methods are becoming more common and more important, but there are not many resources to help beginners get started. People who know Python can use their programming skills to get a head start.
I will present simple programs that demonstrate the concepts of Bayesian statistics, and apply them to a range of example problems. Participants will work hands-on with example code and practice on example problems.
Students should have at least basic Python and basic statistics. If you learned about Bayes’s Theorem and probability distributions at some time, that’s enough, even if you don’t remember it!
Students should bring a laptop with Python 2.x and matplotlib. You can work in any environment; you just need to be able to download a Python program and run it.
Tuesday, January 8, 2013
Are first babies more likely to be late, revisited.
UPDATE: The version of this article with the most recent data is here.
Two years ago I wrote an article called Are first babies more likely to be late?, based on a question that came up when my wife and I were expecting our first child. I compared the pool of first babies to the pool of all other babies, and found:
Two years ago I wrote an article called Are first babies more likely to be late?, based on a question that came up when my wife and I were expecting our first child. I compared the pool of first babies to the pool of all other babies, and found:
- There is a small difference in the mean pregnancy length for the two groups, about 13 hours, but it is not practically or statistically significant.
- If we group babies into Early, On Time, or Late (where On Time is 38, 39 or 40 weeks), first babies are a little more likely to be Early or Late, and less likely to be On Time.
Then yesterday I got the following question from an Unknown correspondent:
While interesting, I can't help but think you need to compare the first and others for the same woman. While may be unlikely it could still be that a tendency exists for a woman's second, third, etc, child comes earlier.
This is an excellent suggestion. It is possible that the variability between people is masking some of the variability between first and later babies. By pairing first and second babies with the same mother, we can control for variation between mothers.
So I ran that experiment, selecting all mothers with at least two children and computing the difference in pregnancy length between the second and first child (so a positive value means the second child was later). Here is the distribution of these value for 4387 women in the NSFG (National Survey of Family Growth):
Visually the distribution looks symmetric, and the summary statistics support that conclusion. The mean is -0.034, which means that (if anything) the second baby is born about 6 hours earlier, but this difference is not statistically significant.
Conclusion: good question, definitely worth running the experiment, but the primary result is the same as what we saw before: no significant difference in the means.
Monday, January 7, 2013
Call for Bayesian case studies
It's been a while since the last post because I have been hard at work on Think Bayes. As always, I have been posting drafts as I go along, so you can read the current version at thinkbayes.com.
I am teaching Computational Bayesian Statistics in the spring, using the draft edition of the book. The students will work on case studies, some of which will be included in the book. And then I hope the book will be published as part of the Think X series (for all X). At least, that's the plan.
In the next couple of weeks, students will be looking for ideas for case studies. An ideal project has at least some of these characteristics:
I am teaching Computational Bayesian Statistics in the spring, using the draft edition of the book. The students will work on case studies, some of which will be included in the book. And then I hope the book will be published as part of the Think X series (for all X). At least, that's the plan.
In the next couple of weeks, students will be looking for ideas for case studies. An ideal project has at least some of these characteristics:
- An interesting real-world application (preferably not a toy problem).
- Data that is either public or can be made available for use in the case study.
- Permission to publish the case study!
- A problem that lends itself to Bayesian analysis, in particular if there is a practical advantage to generating a posterior distribution rather than a point or interval estimate.
Examples in the book include:
- The hockey problem: estimating the rate of goals scored by two hockey teams in order to predict the outcome of a seven-game series.
- The paintball problem, a version of the lighthouse problem. This one verges on being a toy problem, but recasting it in the context of paintball got it over the bar for me.
- The kidney problem. This one is as real as it gets -- it was prompted by a question posted by a cancer patient who needed a statistical estimate of when a tumor formed.
- The unseen species problem: a nice Bayesian solution to a standard problem in ecology.
So far I have a couple of ideas prompted by questions on Reddit:
But I would love to get more ideas. If you have a problem you would like to contribute, let me know!
Subscribe to:
Posts (Atom)