By "Bayes filter", I don't mean spam filtering using a Bayesian classifier, but rather recursive Bayesian estimation, which is used in robotics and other domains to estimate the state of a system that evolves over time, for example, the position of a moving robot. My interest in this topic was prompted by Roger Labbe's book, Kalman and Bayesian Filters in Python, which I am reading with my book club.
The Python code for this article is in this IPython notebook. If you get sick of the math, you might want to jump into the code.
The Bayes filter
A Bayes filter starts with a distribution that represents probabilistic beliefs about the initial position of the robot. It proceeds by alternately executing predict and update steps:- The predict step uses a physical model of the system and its current state to predict the state in the future. For example, given the position and velocity of the robot, you could predict its location at the next time step.
- The update step uses a measurement and a model of measurement error to update beliefs about the state of the system. For example, if we use GPS to measure the location of the robot, the measurement would include some error, and we could characterize the distribution of errors.
The update step uses Bayes theorem, so computationally it involves multiplying the prior distribution by the likelihood distribution and then renormalizing.
The predict step involves adding random variables, like position and velocity, so computationally it involves the convolution of two distributions.
The Convolution Theorem
The Convolution Theorem suggests a symmetry between the predict and update steps, which leads to an efficient algorithm. I present the Convolution Theorem in Chapter 8 of Think DSP. For discrete signals, it says:| DFT(f ∗ g) = DFT(f) · DFT(g) |
That is, if you want to compute the discrete Fourier transform (DFT) of a convolution, you can compute the DFT of the signals separately and then multiply the results. In words, convolution in the time domain corresponds to multiplication in the frequency domain. And conversely, multiplication in the time domain corresponds to convolution in the frequency domain.
In the context of Bayes filters, we are working with spatial distributions rather than temporal signals, but the convolution theorem applies: instead of the "time domain", we have the "space domain", and instead of the DFT, we have the characteristic function.
The characteristic function
This section shows that the characteristic function and the Fourier transform are the same thing. If you already knew that, you can skip to the next section.
If you took mathematical statistics, you might have seen this definition of the characteristic function:
![\varphi_X(t) = \operatorname{E} \left [ e^{itX} \right ]](https://upload.wikimedia.org/math/3/c/8/3c82d2ac41d6cfa6ca018ddbb75c43ae.png)
Where X is a random variable, t is a transform variable that corresponds to frequency (temporal, spatial, or otherwise), and E is the expectation operator.
If you took signals and systems, you saw this definition of the Fourier transform:

where f is a function, 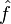 is its Fourier transform, and ξ is a transform variable that corresponds to a frequency.
is its Fourier transform, and ξ is a transform variable that corresponds to a frequency.
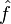 is its Fourier transform, and ξ is a transform variable that corresponds to a frequency.
is its Fourier transform, and ξ is a transform variable that corresponds to a frequency.
The two definitions look different, but if you write out the expectation operator, you get

where p(x) is the probability density function of X, and P(t) is its Fourier transform. The only difference between the characteristic function and the Fourier transform is the sign of the exponent, which is just a convention choice. (The bar above P(t) indicates the complex conjugate, which is there because of the sign of the exponent.)
And that brings us to the whole point of the characteristic function: to add two random variables, you have to convolve their distributions. By the Convolution Theorem, convolution in the space domain corresponds to multiplication in the frequency domain, so you can add random variables like this:
- Compute the characteristic function of both distributions.
- Multiply the characteristic functions elementwise.
- Apply the inverse transform to compute the distribution of the sum.
Back to Bayes
Now we can see the symmetry between the predict and update steps more clearly:
- The predict step involves convolution in the space domain, which corresponds to multiplication in the frequency domain.
- The update step involves multiplication in the space domain, which corresponds to convolution in the frequency domain.
This observation is interesting, at least to me, and possibly useful, because it suggests an efficient algorithm based on the Fast Fourier Transform (FFT). The simple implementation of convolution takes time proportional to N², where N is the number of values in the distributions. Using the FFT, we can get that down to N log N.
Using this algorithm, a complete predict-update step looks like this:
- Compute the FFT of the distributions for position and velocity.
- Multiply them elementwise; the result is the characteristic function of the convolved distributions.
- Compute the inverse FFT of the result, which is the predictive distribution.
- Multiply the predictive distribution by the likelihood and renormalize.
Steps 1 and 3 are N log N; steps 2 and 4 are linear.
I demonstrate this algorithm in this IPython notebook.
For more on the Bayesian part of the Bayes filter, you might want to read Chapter 6 of Think Bayes. For more on the Convolution Theorem, see Chapter 8 of Think DSP. And for more about adding random variables, see Chapter 6 of Think Stats.
Acknowledgements
Thanks to Paul Ruvolo, the other member of my very exclusive reading club, and Wikipedia for the typeset equations I borrowed.
Valuable article for engineer. thanks.
ReplyDelete